Conseils santé, bien être, et développement personnel
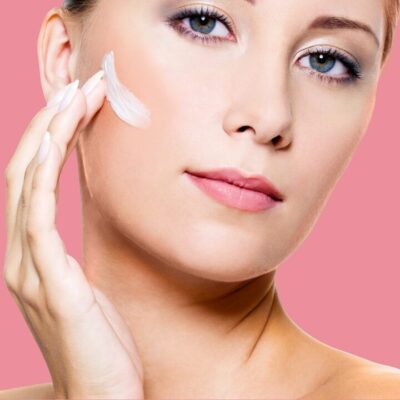
Dans le monde fascinant des produits de beauté et des soins de la peau, l’acide hyaluronique s’impose comme l’acteur étoile des solutions …
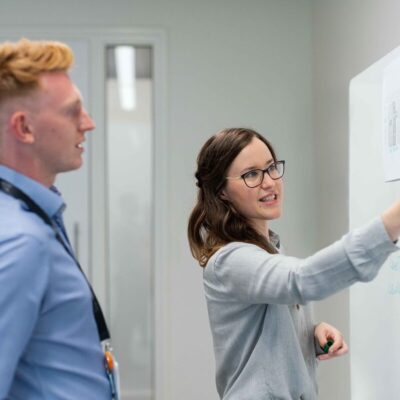
La santé des employés est un enjeu majeur pour les entreprises modernes. Face aux problématiques de stress, de mal-être et de burn-out, …
Bien-être
Découvrez les meilleurs conseils pour favoriser votre bien-être et vivre plus heureux.

Développement personnel
Les meilleurs conseils pour se connaitre et appréhender le monde qui nous entoure, pour bien faire les choses et avancer correctement dans le futur.
Santé
Voici les conseils santé des meilleurs experts, pour une vie à dévorer à pleines dents.
Dernières actualités
Restez au courant des actualités santé comme avec Bioserveur, nutrition, bien-être, nous ne vous laisseront pas perdre ce qu’il faut savoir.